Generation walkthrough for scldm#
This notebook demonstrates single-process data generation using Hydra configs. It mirrors the logic used in our scripts while keeping everything runnable in a notebook. Each section briefly explains the step and then executes the equivalent code.
What you’ll see:
Compose the Hydra config from
experiments/configs/generation.yamlInstantiate the datamodule and model (generator)
Optionally load model weights from a checkpoint
Build a PyTorch Lightning trainer (callbacks/loggers from config)
Run generation and save an
.h5adoutput
Environment and imports#
This cell sets up environment variables and imports the same Python modules used by the script. We keep the imports local to cells to mimic script scoping and to avoid side effects until needed. source
import os
import pathlib
# Match script behavior
os.environ["HYDRA_FULL_ERROR"] = "1"
import hydra
from omegaconf import OmegaConf
from scldm._utils import (
process_generation_output,
process_inference_output,
remap_config,
remap_pickle,
)
# Register useful resolver used in configs
try:
OmegaConf.register_new_resolver("eval", eval)
except ValueError:
# Already registered
pass
print("OmegaConf resolver 'eval' registered.")
OmegaConf resolver 'eval' registered.
Compose the Hydra config#
We compose the configuration directly in-notebook using Hydra’s initialize/compose APIs, pointing to experiments/configs/. Here we load generation.yaml; interpolations and defaults resolve as they would in the scripts.
from hydra import compose, initialize
from omegaconf import OmegaConf
# Initialize Hydra to read from experiments/configs - use relative path
with initialize(config_path="../experiments/configs", version_base="1.2"):
cfg = compose(config_name="generation.yaml")
print(OmegaConf.to_yaml(cfg)[:500]) # preview
paths:
base_data_path: /mnt/czi-sci-ai/project-scg-llm-pvc/release/datasets
base_release_path: /mnt/czi-sci-ai/project-scg-llm-pvc/release/fm_observational
base_experiment_path: /mnt/czi-sci-ai/project-scg-llm-pvc/experiments/release
dataset_paths:
dentate_gyrus:
train: ${paths.base_data_path}/dentategyrus_train.h5ad
test: ${paths.base_data_path}/dentategyrus_test.h5ad
config: ${paths.base_release_path}/dentate_gyrus.yaml
ckpt: ${paths.base_release_path}/denta
Instantiate datamodule and model#
We load the module configuration from cfg.config_file, apply remap_config, and instantiate:
the
DataModuleviahydra.utils.instantiate(cfg.datamodule.datamodule)the model specified in the config (used here for generation)
from omegaconf import OmegaConf
# Module config file path from composed cfg
module_ckpt_path = pathlib.Path(cfg.ckpt_file)
module_config_path = pathlib.Path(cfg.config_file)
print(f"Loading module config from: {module_config_path}")
module_config = OmegaConf.load(module_config_path)
remap_config(module_config)
print("Instantiating datamodule...")
datamodule = hydra.utils.instantiate(cfg.datamodule.datamodule)
print("Instantiating model from config...")
module = hydra.utils.instantiate(module_config.model.module)
print("Datamodule and model ready.")
Loading module config from: /mnt/czi-sci-ai/project-scg-llm-pvc/release/fm_observational/dentate_gyrus.yaml
Instantiating datamodule...
Instantiating model from config...
Datamodule and model ready.
import torch
if module_ckpt_path.exists():
print(f"Loading checkpoint weights from {module_ckpt_path}")
checkpoint = torch.load(module_ckpt_path, map_location="cpu", pickle_module=remap_pickle, weights_only=False)
state_dict = checkpoint["state_dict"] if isinstance(checkpoint, dict) and "state_dict" in checkpoint else checkpoint
module_keys = set(module.state_dict().keys())
filtered_state_dict = {k: v for k, v in state_dict.items() if k in module_keys}
loaded = len(filtered_state_dict)
skipped = len(state_dict) - loaded
missing = len(module_keys) - loaded
if skipped:
print(f"Skipping {skipped} keys not present in module")
if missing:
print(f"Module has {missing} keys not in checkpoint")
print(f"Loading {loaded} matching keys from checkpoint")
module.load_state_dict(filtered_state_dict, strict=False)
else:
print(f"Checkpoint file not found: {module_ckpt_path}")
Loading checkpoint weights from /mnt/czi-sci-ai/project-scg-llm-pvc/release/fm_observational/dentate_gyrus.ckpt
Skipping 154 keys not present in module
Loading 324 matching keys from checkpoint
Build the Trainer, callbacks, and loggers#
We instantiate callbacks and loggers from the training section of the config and construct a PyTorch Lightning Trainer with the same options used for generation runs. Seeding and runtime tweaks match the script behavior.
import pytorch_lightning as pl
import torch
# Match script's runtime setup
torch.set_float32_matmul_precision("high")
torch._dynamo.config.capture_scalar_outputs = True
pl.seed_everything(cfg.seed)
print("Instantiating callbacks and loggers from config...")
callbacks_list = []
for cb_cfg in getattr(cfg.training, "callbacks", {}).values():
callbacks_list.append(hydra.utils.instantiate(cb_cfg, _partial_=False))
loggers_list = []
for lg_cfg in getattr(cfg.training, "logger", {}).values():
loggers_list.append(hydra.utils.instantiate(lg_cfg, _partial_=False))
# Trainer is provided as a partial in config; instantiate callable first
trainer_ctor = hydra.utils.instantiate(cfg.training.trainer) # partial
trainer = trainer_ctor(
devices=1,
strategy="auto",
logger=loggers_list if len(loggers_list) > 0 else False,
callbacks=callbacks_list,
)
# Optional cache tuning, as in script
torch.cuda.empty_cache()
torch._dynamo.config.cache_size_limit = 1000
print("Trainer ready.")
Instantiating callbacks and loggers from config...
Trainer ready.
Run generation and save outputs#
This mirrors the script’s predict flow for generation:
Set the module to eval and call
datamodule.setup("predict")Call
trainer.predict(module, datamodule=datamodule)Post-process with
process_generation_output()to build an.h5adSave results to
cfg.inference_pathwith dataset and type suffix
# Build an override dictionary similar to the script for logging context
# Skip HydraConfig since it's not available in notebook context
override_dict = {"foo": "bar"}
# Predict flow
if cfg.model.module.generation_args is not None or cfg.model.module.inference_args is not None:
module.eval()
datamodule.setup("predict")
output = trainer.predict(module, datamodule=datamodule)
if cfg.model.module.generation_args is not None:
adata = process_generation_output(output, datamodule)
else:
adata = process_inference_output(output, datamodule)
inference_type = "generated" if cfg.model.module.generation_args is not None else "inference"
adata.obs["generation_idx"] = cfg.dataset_generation_idx
else:
print("No generation_args or inference_args specified in the config; skipping predict.")
/mnt/czi-sci-ai/project-scg-llm-pvc/experiments/release/wandb_logger/wandb/run-20251006_144238-r88tof4sINFO Processing generation output
Customization tips#
To switch datasets or models, adjust overrides when composing the config or edit
experiments/configs/generation.yaml.Control behavior via
model.module.generation_args(default here).inference_argscan be used when running inference instead.Output location and naming are controlled by
inference_path,datamodule.dataset, anddataset_generation_idx.
import anndata as ad
import scanpy as sc
adata_test = ad.read_h5ad(pathlib.Path(cfg.datamodule.datamodule.test_adata_path))
adata_test.X = adata_test.layers["X_counts"].copy()
adata_test.obs["dataset"] = "test"
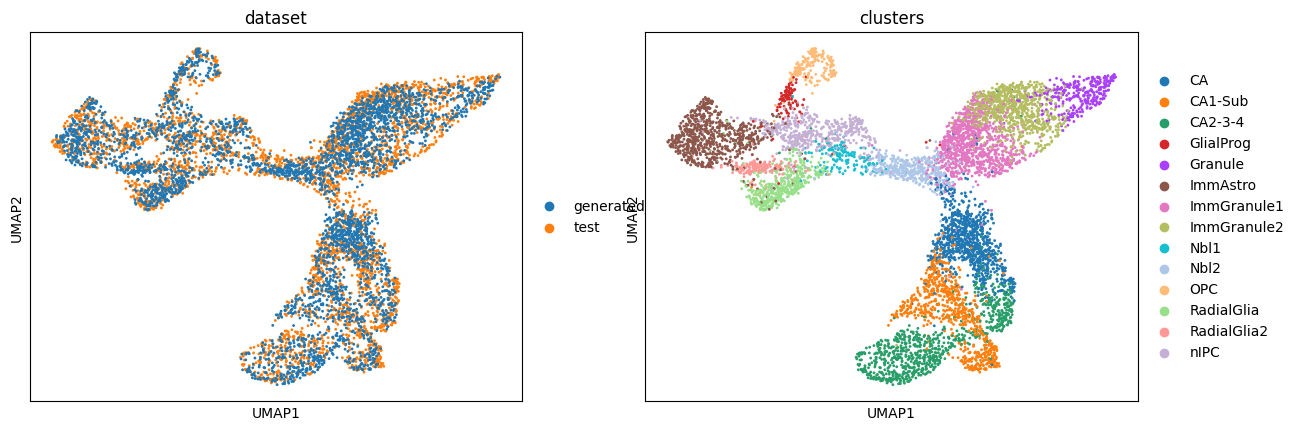
Visualizing generated vs test data#
We compare generated cells with the held-out test set using UMAP to assess overlap and cluster structure.
adata_conditional = ad.concat([adata_test, adata[adata.obs.dataset == "generated_conditional"]], axis=0)
adata_unconditional = ad.concat([adata_test, adata[adata.obs.dataset == "generated_unconditional"]], axis=0)
sc.pp.normalize_total(adata_conditional)
sc.pp.log1p(adata_conditional)
sc.pp.pca(adata_conditional)
sc.pp.neighbors(adata_conditional)
sc.tl.umap(adata_conditional)
sc.pp.normalize_total(adata_unconditional)
sc.pp.log1p(adata_unconditional)
sc.pp.pca(adata_unconditional)
sc.pp.neighbors(adata_unconditional)
sc.tl.umap(adata_unconditional)
sc.pl.umap(
adata_conditional,
color=["dataset", "clusters"],
ncols=2,
)
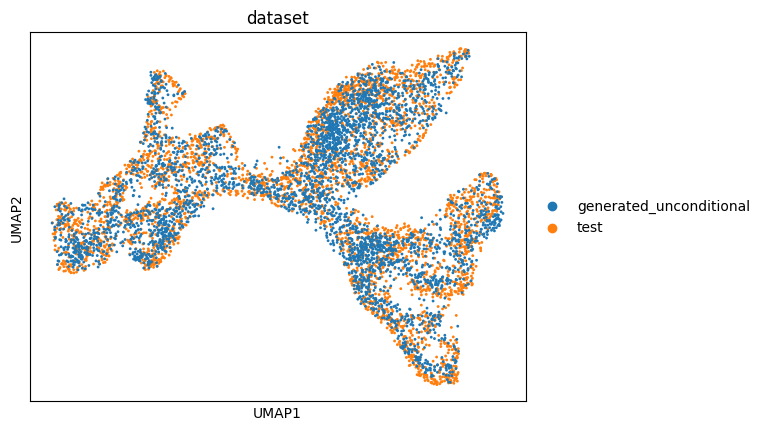
sc.pl.umap(adata_unconditional, color=["dataset"], ncols=2)
Conclusions and next steps#
The notebook demonstrates end-to-end generation using the configured model and datamodule.
Visualizations help assess similarity between generated and test data.
Next steps:
Tune
model.module.generation_args(e.g.,guidance_weight) and re-run.Try different datasets by switching
datamodule.datasetin the config.Save and version results using your preferred logger or artifact store.